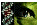
A Molecular Vision Research Article
Isolation of Differentially Expressed Human Fovea Genes: Candidates for Macular Disease
Steven L. Bernstein1*, Diane E. Borst2 and Paul W. Wong1
1National Eye Institute, NIH, Building 6, Rm 337, 9000 Rockville Pike, Bethesda, MD. and 2Department of Anatomy, Uniformed Services University of the Health Sciences, 4301 Jones Bridge Road, Bethesda, MD
*To whom correspondence should be addressed
E-mail: slbernst@umabnet.ab.umd.edu
Current Address:
MSTF 5-65
10 S. Pine St.
University of Maryland School of Medicine
Department of Ophthalmology
Baltimore, MD 21201
ABSTRACT
Purpose: In humans, the fovea is the region of the retina responsible for acute vision. Disorders affecting the fovea are responsible for the majority of cases of untreated blindness in the developed world, yet are poorly understood at the molecular level. Our goal is to identify genes that are preferentially expressed within the human fovea as compared to the midperipheral retina (differential fovea clones).
Materials and Methods: An unamplified fovea cDNA library was differentially screened with cDNA probes derived from either human fovea or midperipheral retina. Rounds of secondary screening and northern analysis were used to verify the expression pattern of a selective number of clones isolated.
Results: Forty-one differential fovea clones were isolated from a screening of 10,000 phage clones (clones). Of these clones, 31.5 % correspond to known sequences present in GenBank/EMBL and 70.7% represent novel human fovea expressed sequence tags (ESTs). Northern analysis of selected clones demonstrated that they represent genes expressed at higher levels in the human fovea than in the midperipheral retina.
Conclusions: Genes that are more highly expressed in the fovea as opposed to the midperipheral retina are likely to represent essential genes for fovea function. Using our fovea cDNA library, we are able to isolate differential human fovea clones at an incidence of 41/10,000 clones screened. We demonstrate that there is a high level of differential gene expression within different regions of the human retina.
INTRODUCTION
The first primary (unamplified) human fovea cDNA library was recently constructed (4, 5). The human fovea is the retinal region responsible for acute (20/20) vision. The fovea has the highest concentration of cone photoreceptors, ganglion cells, and horizontal cells (per mm2), of any retinal region (13) and has not been intensely studied at the molecular level. A great number of disorders and degenerations uniquely affect the human foveo-macular region. These include age-related macular degeneration, cone-rod dystrophies, butterfly macular dystrophy and adult onset foveo-macular dystrophy, among others (9, 22, 27, 28). Disorders of the fovea, and its immediate surrounding area, the macula, are responsible for the majority of cases of untreatable blindness in the developed world, affecting 1 in 25 people over the age of 65 (17).
Genes that are differentially expressed (expressed at higher relative levels) in a given tissue, or tissue region, are likely to be critical to specialized tissue function (16, 25). Changes which alter the expression of these genes or their subsequent gene products may underlie tissue specific diseases. Differential subtraction techniques have proven useful in the identification of a number of retinal expressed/specific genes, including the rd and the rds genes (26, 8, 24). In addition, the methodology of differential cross screening has been used successfully to identify genes that are preferentially expressed in the retina as compared to other tissues (18) as well as to identify genes that are over-expressed in diseased retinas as compared to normal retinas (15). More recently, differential display has been used in an attempt to isolate genes that are differentially expressed in monkey macula as compared to the peripheral retina (7). Because of the potential importance of foveal specific/differentially expressed genes in understanding this histologically distinct region of the retina, we have concentrated on identifying such genes.
Using a differential cross-screening approach, we have selectively isolated clones that are more highly expressed in the fovea than in the peripheral retina from our human fovea cDNA library. We have previously shown by northern and Southern (plaque) analysis that the human fovea cDNA library used in the current study is a non-biased representation of the fovea mRNA population. We chose to use a differential screening approach rather than subtractive cloning, a technique which is inherently biased against the identification of new gene members of established gene families.
MATERIALS AND METHODS
Primary human cDNA phage library
The phage library was constructed by nondirectional ligation of age-normalized human fovea cDNA into lambda zap bacteriophage (5). The fovea is the region, 1.5mm in diameter, centered over the foveola. The titer of this library was 2.0 x 106 PFU/µg cDNA.
RNA
Human donor tissue was obtained from the National Disease Research Interchange (NDRI, Philadelphia, PA). Foveae and midperipheral retinal tissue was dissected from 10 pairs of donor eyes ranging from 2-79 years of age. Total RNA was extracted using RNAzol B (Tel-Test, Friendswood, TX). Total human liver and brain RNA was obtained from Clontech (Palo Alta, CA).
Northern Blots
Five µg of total RNA per lane, was denatured and electrophoresed on 1.25% formaldehyde gels. Following electrophoresis, the RNA was transferred and immobilized on Genescreen plus membranes (Dupont; Boston, MA).
Preparation of the mixed cDNA probes from human fovea and midperipheral retina
Poly (A+)RNA from both preparations were purified from the human fovea and midperipheral retina (Oligotex; Qiagen, Chatsworth, CA). First strand cDNA was synthesized from one µg of poly (A+) RNA from each preparation, using the 3' RACE system (Life Technologies, Gaithersburg, MD). Single stranded cDNA was tailed with poly (dC) using terminal transferase. Second strand synthesis was performed using the 5' anchor RACE primer (Life Technologies). These mixed cDNA probes were amplified, using the universal amplification primer (UAP; Life Technologies), via the polymerase chain reaction (PCR).
Radiolabeling and hybridization of probes
Mixed cDNA probes used for differential cross screening were random-primed labeled using a combined mixture of 32P dCTP, 32P dATP and 32P dGTP in a standard labeling reaction (Prime-It II, Stratagene, La Jolla, CA). Specific inserts from selected clones were labeled with 32P dCTP in a standard random priming labeling reaction. Minimum activity used per hybridization was 5 x 106 CPM/ml of hybridization buffer (Hybrisol II, Oncor, Gaithersburg, MD). Northern blots/plaque lifts were prehybridized in Hybrisol II for 3 hours at 63°C and incubated with the probe in hybridization buffer for 18 hours at 63°C. Following hybridization, blots were stringently washed in 0.2X SSC at 63°C and exposed for 1-3 days at -70°C to Kodak X-O-Mat film.
Isolation of differentially expressed clones
Duplicate plaque lifts were made from 150mm plates with 6 X 102 PFU of the library/plate. One set of plaque lifts was screened with a mixed cDNA human fovea probe. The duplicate set of plaque lifts was screened with a mixed cDNA human midperipheral probe. Candidate clones for genes that are more highly expressed in the fovea were identified by precise alignment of both autoradiographs of the duplicate lifts reacted with the differential screening probes, with the plate containing the original plaques. Clones were isolated and placed in buffer containing 100mM NaCl/50mM MgCl2 pH 7.5 buffer.
Secondary screening
An aliquot of each eluted recombinant phage clone was amplified using PCR, with T3 and T7 RNA polymerase primers flanking the cDNA insert site. An aliquot of the PCR product was spotted onto marked, duplicate nylon filter arrays. These filter arrays (secondary blots) were hybridized with either the radiolabeled (a) foveal cDNA probe or (b) midperipheral retina probe. Secondary blots were then stringently washed in 0.2XSSC at 63°C and exposed to Kodak X-AR film.
Removal of mitochondrial sequences
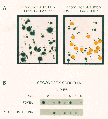
In order to eliminate the bulk of clones that represent redundant mitochondrial messages from our selected pool of clones, we reacted one set of secondary blots with a mixed mitochondrial DNA probe, made with primers specific for the mitochondrial genome (
Fig. 2A & B). Amplified mitochondrial DNA was labeled by ECL Direct (Amersham Corporation, Arlington Heights, IL). The blots were incubated with the probe and then washed stringently (6M urea/0.5XSSC at 45°C). The hybridized probe was detected by a chemiluminescent methodology using the manufacturers recommended protocols.
Sequencing and sequence analysis
Clones were sequenced using fluorescent-labeled dideoxynucleotide technology (Prism; Applied Biosystems, Foster City, CA). The sequenced cDNA inserts were run on an ABI model 370A automated sequencer. Sequence information was analyzed and compared to the GenBank/EMBL (6/95) databases using the FASTA program available through the GCG package of programs (version 8.0). Similar sequences were defined as >55% homology over 70bp or greater. Identical sequences were defined as >95% identity over the entire length of the sequence.
RESULTS
Differential screening of the human fovea library
 10,000 clones were differentially screened. A representative set of autoradiographs of duplicate plaque lifts are shown in
Fig. 1A. Lifts were probed with either the 32P-labeled fovea cDNA (on the left), or midperipheral retina cDNA probe (on the right). A total of 411 clones that showed higher levels of hybridization with the fovea versus the midperipheral probe were isolated. 220 of these clones proved to represent putative differential fovea sequences after secondary screening (Fig. 1B).
10,000 clones were differentially screened. A representative set of autoradiographs of duplicate plaque lifts are shown in
Fig. 1A. Lifts were probed with either the 32P-labeled fovea cDNA (on the left), or midperipheral retina cDNA probe (on the right). A total of 411 clones that showed higher levels of hybridization with the fovea versus the midperipheral probe were isolated. 220 of these clones proved to represent putative differential fovea sequences after secondary screening (Fig. 1B).
Identification of mitochondrial sequences
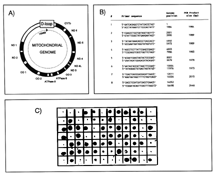 Clones coding for mitochondrial sequences are highly abundant in the fovea library, making up nearly 20% of all clones (4). We have previously shown by northern analysis that different mitochondrial genes appear to be differentially expressed in the fovea as compared to the peripheral retina, although it should be noted that mitochondrial gene expression is extremely high in both the fovea and the peripheral retina (4). Initial sequencing of isolated fovea differentially expressed clones indicated that a significant percentage of these clones contained mitochondrial sequences (data not shown). The mitochondrial sequence screen of the secondary blots was performed to minimize the re-sequencing of these redundant sequences. Sequencing of 36 mitochondrial probe positive clones revealed that 95% of clones isolated by this method showed identity to mitochondrial messages (data not shown). In total, 124 putative fovea enriched clones on the secondary blots showed strong signals when screened with the mixed mitochondrial probe (Fig. 2C). These clones were eliminated from subsequent analysis. Clones generating no, or weak, signals with the mitochondrial probe were included for further analysis (Fig. 2C).
Clones coding for mitochondrial sequences are highly abundant in the fovea library, making up nearly 20% of all clones (4). We have previously shown by northern analysis that different mitochondrial genes appear to be differentially expressed in the fovea as compared to the peripheral retina, although it should be noted that mitochondrial gene expression is extremely high in both the fovea and the peripheral retina (4). Initial sequencing of isolated fovea differentially expressed clones indicated that a significant percentage of these clones contained mitochondrial sequences (data not shown). The mitochondrial sequence screen of the secondary blots was performed to minimize the re-sequencing of these redundant sequences. Sequencing of 36 mitochondrial probe positive clones revealed that 95% of clones isolated by this method showed identity to mitochondrial messages (data not shown). In total, 124 putative fovea enriched clones on the secondary blots showed strong signals when screened with the mixed mitochondrial probe (Fig. 2C). These clones were eliminated from subsequent analysis. Clones generating no, or weak, signals with the mitochondrial probe were included for further analysis (Fig. 2C).
Sequence analysis of differentially selected fovea clones
Ninety-six clones remained after screening with the mixed mitochondrial probe. Sixty-five clones yielded informative sequence which was compared to sequences in the GenBank/EMBL database. Of these 24 were mitochondrial in origin and discarded from further analysis. Remaining clones showing identity or similarities to sequences in these databases, other than mitochondrial in nature, are shown in
Table 1. Known gene sequences such as calmodulin, kv2.1 potassium channel, and the EST clone 79E10 are represented a number of times in this list, suggesting that there has been a selective isolation of specific clones in our differential cross screen.
 In total, 12 clones (31.5%; 12/41) showed putative identity to known GenBank/EMBL entries (Fig. 3). Twenty-nine clones (70.7%, 29/41) represent new ESTs. Twenty-four of these clones (58.5%; 12/41) show some sequence similarity to known sequences in GenBank/EMBL and are likely to represent new members of previously identified gene families (Fig. 3). Five clones (12.2%; 5/41) represent unique, previously unidentified, differentially expressed foveal gene sequences (Fig. 3).
In total, 12 clones (31.5%; 12/41) showed putative identity to known GenBank/EMBL entries (Fig. 3). Twenty-nine clones (70.7%, 29/41) represent new ESTs. Twenty-four of these clones (58.5%; 12/41) show some sequence similarity to known sequences in GenBank/EMBL and are likely to represent new members of previously identified gene families (Fig. 3). Five clones (12.2%; 5/41) represent unique, previously unidentified, differentially expressed foveal gene sequences (Fig. 3).
Northern analysis of differentially selected foveal sequences
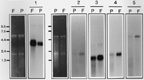 Northern hybridization analysis remains the standard method to determine messenger abundance in a cellular messenger population. Northern analysis has been performed on 20 of the putative fovea differential clones, 19 of which have shown higher levels of expression in the fovea than midperipheral retina. Representative results are shown in Fig. 4. Included in Fig. 4 is the northern analysis of retinal RNA samples with NADH4, a mitochondrial gene that is known to be more highly expressed in fovea as compared to midperipheral retina (5), clone HFD122210 (calmodulin), clone HFD122215 (similar to rat aldolase C), and clones HFD010910 and HFD121219 (novel foveal ESTs). In all cases, genes analyzed show a higher level of expression in the fovea as compared to the midperipheral retina.
Northern hybridization analysis remains the standard method to determine messenger abundance in a cellular messenger population. Northern analysis has been performed on 20 of the putative fovea differential clones, 19 of which have shown higher levels of expression in the fovea than midperipheral retina. Representative results are shown in Fig. 4. Included in Fig. 4 is the northern analysis of retinal RNA samples with NADH4, a mitochondrial gene that is known to be more highly expressed in fovea as compared to midperipheral retina (5), clone HFD122210 (calmodulin), clone HFD122215 (similar to rat aldolase C), and clones HFD010910 and HFD121219 (novel foveal ESTs). In all cases, genes analyzed show a higher level of expression in the fovea as compared to the midperipheral retina.
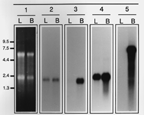 We also compared expression of these messages in human brain and liver total RNA (
Fig. 5). All probes examined (NADH4, HFD122215, HFD010910, and HFD121219) generated a much stronger signal in RNA from brain than from liver. Clones HFD122215 and HFD121219 show very little hybridization to liver RNA and may therefore represent neural-specific (enriched) gene sequences.
We also compared expression of these messages in human brain and liver total RNA (
Fig. 5). All probes examined (NADH4, HFD122215, HFD010910, and HFD121219) generated a much stronger signal in RNA from brain than from liver. Clones HFD122215 and HFD121219 show very little hybridization to liver RNA and may therefore represent neural-specific (enriched) gene sequences.
DISCUSSION
The current study represents the first direct mass screening for clones that are differentially expressed in the human fovea. Previously, few fovea differentially expressed genes have been identified (7, 21). In the current study we have identified 41 clones representing putatively differential fovea expressed sequences. Twenty-nine of these clones represent new gene discoveries. Clones corresponding to new genes with nucleotide sequence similarities to sequences in GenBank/EMBL make up the largest group of single copy gene sequences (24/41 clones or 58.5%). Five of 41 clones (12.2%) represent new human gene sequences with no similarities to known sequences. Of the differential fovea clones tested by northern analysis, all have shown detectable hybridization to foveal RNA and therefore represent middle abundant to abundant expressed sequences in the foveal mRNA sequence pool.
Of the clones isolated, 12 have sequence identity with 6 known gene sequences (calmodulin, Kv2.1 Shab potassium channel, protein kinase, creatine kinase ß-subunit, glutathione peroxidase, and clathrin heavy chain). A common feature of these 6 genes is that they all represent known human brain ESTs. In the northern analyses shown in Figs. 4 and 5, all of the differential selected fovea clones tested show an apparent neural tissue specificity, with levels of hybridization higher in fovea, peripheral retina, and brain than in liver.
EST analysis of cDNA libraries has been useful in determining unique patterns of expression in different organs, with the frequency of specific genes or families of related genes identified being used as an index for biological activity or function (1, 2). Of the known gene sequences with identical matches to differential fovea ESTs defined in the present study, two were detected in the differentially selected clones a multiple number of times: Gene sequences for calmodulin were detected five times (HFD121234, HFD121235, HFD122210, HFD122221, AND HFD122224) and kv2.1 Shab potassium channel protein were detected three times (HFD072011, HFD121220, and HFD122205). These findings suggest that the relative abundance of these messages in the fovea is high and that these genes may underlie relevant biological functions in the fovea.
Calmodulin is a ubiquitous eukaryotic calcium binding protein. In neurons, calcium binds to calmodulin, and, in turn, this complex can bind to other proteins and alter their activities (10). In vertebrate rod photoreceptor cells, Ca2+-calmodulin can bind to the cGMP-gated cation channel, which underlies the phototransduction cascade, and reduce the affinity of cGMP for the channel (14). Modulation of the association of the Ca2+-calmodulin complex with the channel effects photorecovery and light adaptation. The cone photoreceptor cGMP-gated channel that underlies phototransduction in these cells is genetically distinct from that found in the rod photoreceptor (6). However, although the majority of proteins along the phototransduction pathway enlists different isoforms in rods and cones, the overall mechanism of action is similar in both cells types. Therefore, detection of calmodulin gene expression in the fovea is not surprising. The higher levels of calmodulin gene expression in fovea as compared to the peripheral retina reflects a biological difference between the two regions. Whether this reflects a physiological difference in calmodulin action in cone cell phototransduction versus rod cell phototransduction is not known. In rod photoreceptors there are additional proteins that can mimic the effects of calmodulin (29). Thus, if the relative role of calmodulin in cone phototransduction is important and there is a genetic or functional defect which compromises its function it may lead to a cone photoreceptor cell specific dysfunction. The calmodulin-1 gene locus has been localized to chromosome 14q24-q31 (3). In addition, rod monochromacy, a recessive disorder in which cones are present but functionally defective, has also been mapped to human chromosome 14 (23).
Potassium channels are a family of membrane proteins present in all eukaryotic cells. Their diverse functions include maintaining membrane potential, regulating cell volume and modulating electrical excitability in neurons (19). Four sequence-related potassium channel genes, Shaker , Shaw , Shab and Shal , have been identified in Drosophila (19). Each member of this Drosophila gene family has been shown to have a human homologue. Two human Shaw -related genes, kv3.2 (KCNC2) and kv3.3 (KCNC3), which have been localized to chromosome 19q13.3-q13.4 (12), are currently being considered as candidate genes for Chromosome 19 cone-rod dystrophy (11). In our current study, we isolated three independent clones for human kv2.1 (KCNB1), a Shab potassium channel protein gene. This represents the first report of Shab potassium channel protein gene expression in the retina. The multiple isolation of this gene sequence from the fovea library suggests that kv2.1 may play an important role in foveal homeostasis. The kv2.1 gene has been recently localized to chromosome 20q13.2 (20), and because of its known foveal expression, should be examined as a candidate for genetic foveal/macular disorders that map to this region of the genome.
Our study has identified a significant number of new genes that are differentially expressed in the human fovea. In addition to gene discovery, identifying regionally differentially expressed genes is important for understanding regional retinal function. Identification of differentially expressed foveal genes coupled with their localization in the human genome will be useful in determining potential associations with genetically mapped foveal/macular disease loci. The construction of a differentially selected fovea expressed gene 'bank' provides a much needed resource for accelerating the elucidation of the molecular lesions underlying different inherited foveo-macular retinopathies.
ACKNOWLEGEMENTS
We thank Dr. Radha Ayyagari (National Eye Institute [NEI], National Institutes of Health [NIH], Bethesda, MD) for helpful discussions, Dr. Claire Marcus Bernstein (Bethesda, MD) for helpful discussions and review of the manuscript, Dr. Gerald J. Chader (NEI, NIH, Bethesda, MD), and Dr. Carl Kupfer, Director, NEI for his support. P.W thanks the RP Research Foundation (Canada) for their support.
REFERENCES
1. Adams, M.D., Kerlavage, A.R., Fields, C., and Venter, J.C. 3400 new expressed sequence tags identify diversity of transcripts in human brain. Nat. Genet. 1993; 4, 256
267. 
2. Adams, M.D., Soares, M.B., Kerlavage, A.R., Fields, C., and Venter, J.C. Rapid cDNA sequencing (expressed sequence tags) from a directionally cloned human infant brain cDNA library. Nat. Genet. 1993; 4, 373-380. 
3. Berchtold, M.W., Egli, R., Rhyner, J.A., Hameister, H. and Strehler, E.E. Localization of the human bona fide calmodulin genes CALM1, CALM2, and CALM3 to chromosomes 14q24-q31, 2p21.1-p21.3, and 19q13.2-q13.3. Genomics. 1993; 16, 461-465. 
4. Bernstein, S.L., Borst, D.E., Neuder, M.E. and Wong, P.W. Characterization of a human foveal primary cDNA library and regional differential gene expression in the human retina. ms submitted.
5. Bernstein, S.L., Borst, D.E. and Wong, P.W. Characterization of a human foveal primary cDNA library and isolation of candidate clones for human macular dystrophies. Invest. Ophthal. Vis. Sci. 1995; 36, (4) 2860, abstract.
6. Bonigk, W., Altenhofen, W., Muller, F., Dose, A., Illing, M., Molday, R.S., Kaupp, U.B. Rod and cone photoreceptor cells express distinct genes for cGMP-gated channels. Neuron. 1993; 10, 865-877. 
7. Bowes, C.A. and Hageman, G.S. Differential display and identification of genes differentially expressed in the primate macula Invest. Ophthal. Vis. Sci. 1995; 36, (4) 2859, abstract.
8. Bowes, C., Danciger, M., Kozak, C.A., and Farber, D.B. Isolation of a candidate cDNA for the gene causing retinal degeneration in the rd mouse. Proc. Natl. Acad. Sci. USA. 1989; 86, 9722-9726.
9. Gass, J.D.M. Atlas of Macular Diseases. St. Louis: Mosby; 1987: 2nd ed.,
10. Ghosh, A. and Greenberg, M.E. Calcium signaling in neurons: Molecular mechanisms and cellular consequences. Science. 1995; 268, 239-47. 
11. Gregory, C.Y., Evans, K., Bird, A.C. and Bhattacharya, S.S. Linkage refinement and candidate gene screening for chromosome 19q cone-rod retinal dystrophy. Invest. Ophthal. Vis. Sci. 1995; 36, (4) 4098, abstract.
12. Haas, M., Ward, D., Lee, J., Roses, A.D., Clarke, V., D'Eustachio, P., Lau, D., Vega-Saenz de Miera, E., Rudy, B. Localization of Shaw-related K+ channel genes on mouse and human chromosomes. Mammalian Genome. 1993; 4, 711-5. 
13. Hendrickson, A.E. Primate foveal development: A microcosm of current questions in neurobiology. Invest. Ophth. Vis. Sci. 1994; 35, 3129-3133.
14. Hsu, Y-T. and Molday, R.S. Modulation of the cGMP-gated channel of rod photoreceptor cells by calmodulin. Nature. 1993; 361, 76-79. 
15. Jones, S.E., Meerabux, J.M.A., Yeats, D.A., and Neal, M.J. Analysis of differentially expressed genes in retinitis pigmentosa retinas altered expression of clusterin mRNA. FEBS. 1992; 300, 279-282. 
16. Liew, C.C., Hwang, D.M., Fung, Y.W., Laurenssen, C., Cukerman, E., Tsui, S. and Lee, C.Y. A catalogue of genes in the cardiovascular system as identified by expressed sequence tags. Proc. Natl. Acad. Sci, U.S.A., 1994; 91, 10645-10649. 
17. MacDonald, H.R., Schatz, H., Johnson, R.N. and Madeira, D. Acquired macular degeneration. In: Duanes Textbook of Ophthalmology. Philadelphia: J.B. Lippencott; 1990: V.3, 23. W. Tasman, ed.
18. McInnes, R.R., Manara, S., Sexsmith, E., Warsh, J., Sadler, S., and Taylor, R.G. The isolation of abundant retinal-specific cDNAs: candidate genes for retinal diseases and probes for the study of retinal development. Am. J. Hum. Genet. 1988; 43, A195, abstract.
19. McPherson, J.D., Wasmuth, J.J., Chandy, K.G., Swanson, R., Dethlefs, B., Chandy, G., Wymore, R., Ghanshani, S. Chromosomal localization of 7 potassium channel genes. Cytogenet. Cell Genet. 1991; 58, 1979.
20. Melis, R., Stauffer, D., Zhao, X., Zhu, X.L., Albrecht, B., Pongs, O., Brothman, A., and Leppert, M. Physical and genetic localization of a Shab subfamily potassium channel (KCNB1) gene to chromosomal region 20q13.2. Genomics. 1995; 25, 285-87. 
21. Nathans, J., Thomas, D., and Hogness, D.S. The molecular genetics of color vision: The genes encoding blue, red and green pigments. Science. 1986; 232, 193-202. 
22. Nichols, B.E., Sheffield, V.C., Vandenburgh, K., Drack, A.V., Kimura, A.E. and Stone, E.M. Butterfly-shaped pigment dystrophy of the fovea caused by a point mutation in codon 167 of the rds gene. Nature Genetics. 1993; 3, 202-207. 
23. Pentao, L.. Lewis, R.A., Ledbetter, D.H., Patel, P.I. and Lupski, J.R. Maternal uniparental isodisomy of chromosome 14: association with autosomal recessive rod monochromacy. Am. J. Hum. Genet. 1992; 50, 690-699. 
24. Swaroop, A., Xu, J.Z., Agarwal, N. and Weissman, S.M. A simple and efficient cDNA library subtraction procedure: Isolation of human retina-specific cDNA clones. Nucleic Acids Res. 1991; 19, 1954. 
25. Takeda, J., Yano, H., Eng, S., Zeng, Y. and Bell, G.I. A molecular inventory of human pancreatic islets: sequence analysis of 1000 cDNA clones. Hum. Mol. Genet. 1993; 2, 1793-1798. 
26. Travis, G., Brennan, M., Danielson, P., Kozak, C., and Sutcliffe, J. Identification of a photoreceptor-specific mRNA encoded by the gene responsible for retinal degeneration slow (rds). Nature. 1989; 338, 70-73.
27. Weber, B.H.F., Vogt, G., Pruett, R.C., Stohr, H. and Felbor, U. Mutations in the tissue inhibitor of metalloproteinase 3 (TIMP-3) in patients with Sorsby's fundus dystrophy. Nature Genetics. 1994; 8, 352-356. 
28. Wells, J., Wroblewski, J., Keen, J., Inglehearn, C., Jubb, C., Eckstein, A., Jay, M., Arden, G., Bhattacharya, S. and Bird, A.F. Mutations in the human retinal degeneration slow (RDS) gene can cause either retinitis pigmentosa or macular dystrophy. Nature Genetics. 1993; 3, 213-218. 
29. Yau, K-W. Phototransduction mechanism in retinal rods and cones. Invest. Ophthal. Vis. Sci. 1994; 35, 9-32.
 Return to Molecular Vision homepage
Return to Molecular Vision homepage
Received 2 October 1995 | Revised 28 November 1995 | Accepted 15 December 1995 | Uploaded 21 December 1995
Referencing Note: This article may be referenced as: Mol. Vis. 1:4, 1995.
Alternatively, this article may be referenced by its unique URL:
http://www.emory.edu/molvis/v1/bernstein
© 1995 Molecular Vision  10,000 clones were differentially screened. A representative set of autoradiographs of duplicate plaque lifts are shown in
Fig. 1A. Lifts were probed with either the 32P-labeled fovea cDNA (on the left), or midperipheral retina cDNA probe (on the right). A total of 411 clones that showed higher levels of hybridization with the fovea versus the midperipheral probe were isolated. 220 of these clones proved to represent putative differential fovea sequences after secondary screening (Fig. 1B).
10,000 clones were differentially screened. A representative set of autoradiographs of duplicate plaque lifts are shown in
Fig. 1A. Lifts were probed with either the 32P-labeled fovea cDNA (on the left), or midperipheral retina cDNA probe (on the right). A total of 411 clones that showed higher levels of hybridization with the fovea versus the midperipheral probe were isolated. 220 of these clones proved to represent putative differential fovea sequences after secondary screening (Fig. 1B).