![]() Appendix 1 of
Hamer, Mol Vis 2000;
6:265-286.
Appendix 1 of
Hamer, Mol Vis 2000;
6:265-286.
Appendix 1. Equation Derivations
A. Derivation of PDE activation, including an R*<--R Back-Reaction Without Ca++-Feedback Onto R* lifetime
Rhodopsin activation and depletion are treated as first-order process [17,19-21],
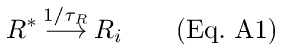
with a corresponding differential equation,
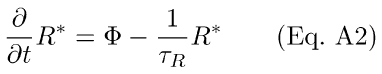
Here Φrepresents the number of photoisomerizations elicited by a brief flash of light.
An ordinary differential equation for PDE; activation may be derived from a simplified enzymatic reaction [19,20]:
The quantity E represents the number of molecules of a complex of transducin (T) and PDE, [T·PDE] = E, which is activated enzymatically in a single step by R* with rate K.
The corresponding differential equation describing the rate of change of activated E* is given by Eq. A4
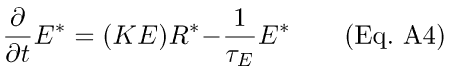
This scheme assumes that the depletion of E* is a first-order process with rate constant 1/τE. If it is also assumed that the above reaction is not stoichiometrically limited by the amount of E available, then the coefficient governing the accretion of E*, KE, may be approximated as a pseudo-first-order rate constant, νrp, yielding
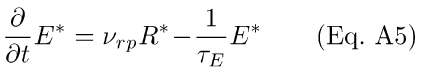
The scheme depicted in Eqs. A1 through A5 is not, strictly speaking, correct. The actual chemistry involves a number of additional steps, including enzymatic activation of the G-protein, or transducin (T). The full PDE activation process may consist of 6-8 "micro-steps" [21,66]. Nevertheless, over the past few years, it has been proposed that the PDE activation process is well approximated by summarizing all the reactions (the internal dynamics of which are, for the most part, unknown) as a two-stage process.
Introduction of R*<--Ri back-reaction
It has been observed in several species that the response to a prolonged step of light has multi-phasic dynamics at step-offset (early fast recovery, followed by a much slower recovery with some occasional resonant behavior in the transition; toad: [67]; salamander: [22,28]; monkey: [18]). One mechanism that can reproduce this behavior is the inclusion of a back-reaction from Ri to R* [17]. Recently, Hamer & Tyler [68] and Hamer [15,69,70] incorporated this back-reaction in computational phototransduction models and were able to capture a number of salient features of amphibian rod responses, including the acceleration of the Tsat versus ln(I) function at high intensities [12] and the multi-phasic step-offset response. The differential equations for accretion and deletion of R* and Ri may be derived from the chemical scheme in Eqs. A6a through A6b (equivalent to scheme by Forti et al. [17]). It is worth noting that this reaction scheme is functionally equivalent to other schemes in which additional forms of inactivated rhodopsin (apoprotein) may continue to activate PDE at a slow rate [71-73].
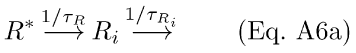

These may be written in differential equation form as
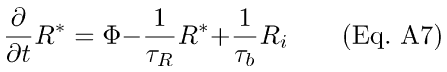
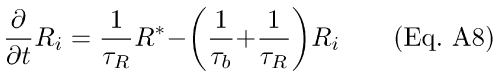
B. Derivation of Ca++-Feedback Onto R* lifetime Via Rec/RK in the Presence of an R*<--R Back-Reaction
The RecRK model adds Ca-sensitivity to the inactivation of R*. The first reaction is the cooperative binding of Ca++ by Rec:
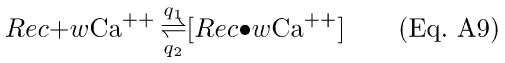
The complex [Rec·wCa++] will be symbolized by Rec*.
The second reaction is a reversible disinhibition of RK to its active form (RK*) by interaction with Rec*:
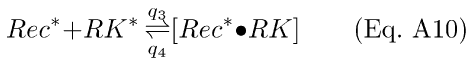
where the complex [Rec*·RK] is the inhibited form of RK. The first reaction (Eq. A9) leads to a differential equation for Rec*:
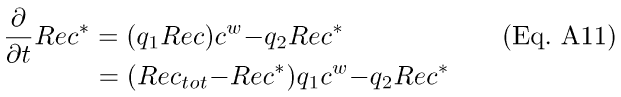
where Rectot = Rec* + Rec, the total amount of Rec in the rod. It was assumed that this reaction reached equilibrium rapidly, such that the formation of Rec* follows the instantaneous level of internal Ca++. Setting Eq. A11 equal to zero and solving for Rec*, we get the steady-state equation
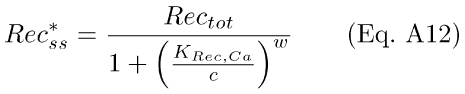
where KRec,Ca = (q2/q1)1/w, the Ca++ concentration at which half of Rec is bound to Ca++.
The second reaction (Eq. A10), the disinhibition of RK, can be written in differential equation form as
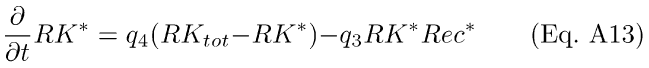
RKtot is the total amount of RK in the cell, and (RKtot - RK*) is the amount of inactive RK.
Eq. A13 has the steady-state solution,
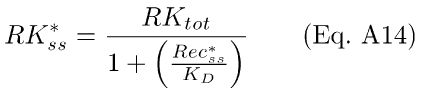
where KD = q4/q3, the concentration of Rec* corresponding to 1/2-activation of RK.
For simplicity, it was assumed that RK* leads to R* phosphorylation (-inactivation) in two steps. Step 1 is the interaction between R* and the disinhibited (activated) RK (=RK*) to form a complex, [R*·RK*]
Step 2 is incorporation of a number of phosphates (P) onto the [R*·RK*] [74], yielding a new complex [nP·R*] and releasing free RK*. The complex [nP·R*] is treated as inactivated rhodopsin (Ri)
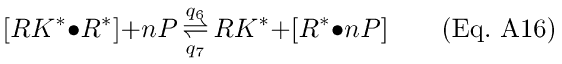
Only Step 2 is assumed to be reversible, and q5 is assumed to be rate-limiting in the two forward reactions.
Capping by arrestin, the final mechanism of R* inactivation, is not explicitly modeled [74], but is assumed to be simultaneous with, and stoichiometrically equivalent to the phosphorylation process.
Under these assumptions, the following differential equations result:
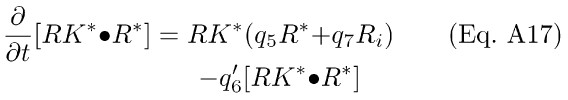
where q'6 = q6Pn, and Pn is a constant.
Under the assumption that reaction A15 is rate-limiting, phosphorylation will proceed at a rate governed by the instantaneous level of the complex [RK*·R*], and a steady-state solution to Eq. A17 may be used:
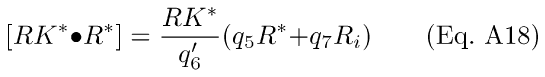
The differential Eqs. for R* and Ri dynamics may now be written as:
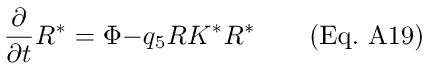
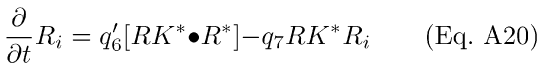
Substituting Eq A18 into Eq A20, we get
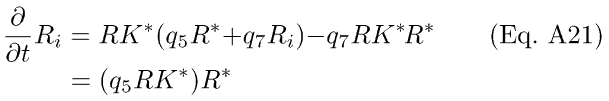
In Eq. A19, the Ca++-sensitivity is expressed in the modulation of R* lifetime by RK*.
In addition to decay of R* to Ri (with rate-constant 1/τR; Eq. A6a), depletion of Ri and the back-reaction to R* from Ri are assumed to occur by separate pathways.
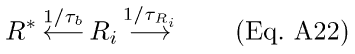
The back-reaction to R* proceeds with rate 1/τb, and Ri is depleted with rate 1/τRi. The resulting differential equations for R* and Ri with Ca++-modulation by RK are analogous to Eqs. A7-8:
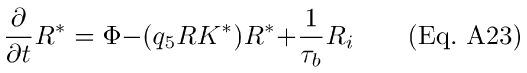
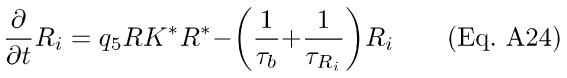
The quantity (q5RK*) in Eq. A23 corresponds to the R* decay time constant, (1/τR) in Eqs. A2, A7-8. However, in Eq. A23, the rate of decay of R* is a time-dependent function of Ca++. In the presence of light, Ca++ decreases, leading to a decrease in the steady-state amount of Rec* (Eq. A12). A decrease in Rec* leads, in turn, to an increase in the amount of RK* (Eq. A14), which translates to an effective increase in the rate-constant governing the depletion of R* (Eq. A23).