![]() Figure 5 of
Bernstein, Mol Vis 1998;
4:24.
Figure 5 of
Bernstein, Mol Vis 1998;
4:24.
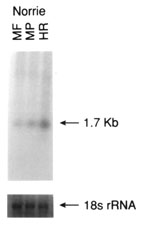
Figure 5. Northern analysis of total RNA using cDNA probes from non retina-specific genes known to cause retinal disease
Total RNA (5 mg) from each of pooled monkey fovea (MF), pooled monkey midperipheral retina (MP), monkey mixed choroid/RPE tissue (MR/Ch), human total retina (HR), pooled human fovea (HF), pooled human midperipheral retina (HP), human RPE/choroid complex (HR/Ch), human ciliary body (HCB), human brain (HB), and human liver (HL) were denatured, electrophoresed, transferred to nylon membranes and reacted with the radiolabelled cDNA probes as described in methods. Following hybridization, blots were stringently washed (63 °C/0.2X SSC), and exposed to X-ray film at -70 °C for varying lengths of time. Shown below each autoradiograph are the ethidium bromide stained 18S ribosomal RNA bands from the original agarose gel from which the analyzed blot was made. The ethidium bromide stained 18S rRNA band was analyzed densitometrically and used as an internal RNA loading standard.
Figure 5A. CHM (choroideremia), 14 day exposure
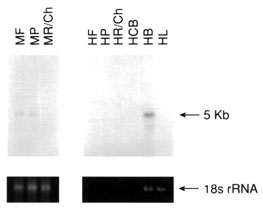
Figure 5B. Ornithine aminotransferase, 12 day exposure
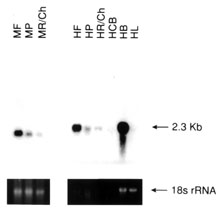
Figure 5C. NDP (Norrie disease protein), 1 day exposure