Figure 3. Comparison and reevaluation of variants in VSX1. A: Distribution and frequency of variants in VSX1 in gnomAD and our cohort, and variants reported in literature. The scoring criteria of prediction are as follows: the total
score is 25 points. For SIFT, a rating of D scores 5 points, and T scores 0 points. For PolyPhen-2, a rating of D, PD and
T scores 5, 3 and 0 points, respectively. For PROVEAN, D scores 5 points and N scores 0 points. For REVEL/ CADD, a score above
the 95% cutoff scores 5 points, between the 75% and 95% cutoffs scores 3 points, and below the 75% cutoff scores 0 points.
In addition, each occurrence of gnomAD Allele count deducts 1 point. B: Rate of types of variants in VSX1 in gnomAD, our cohort and reported studies. CI=Confidence interval. (C) The sankey diagram of reevaluation of reported variants
in VSX1 from HGMD or Clinvar to this study.
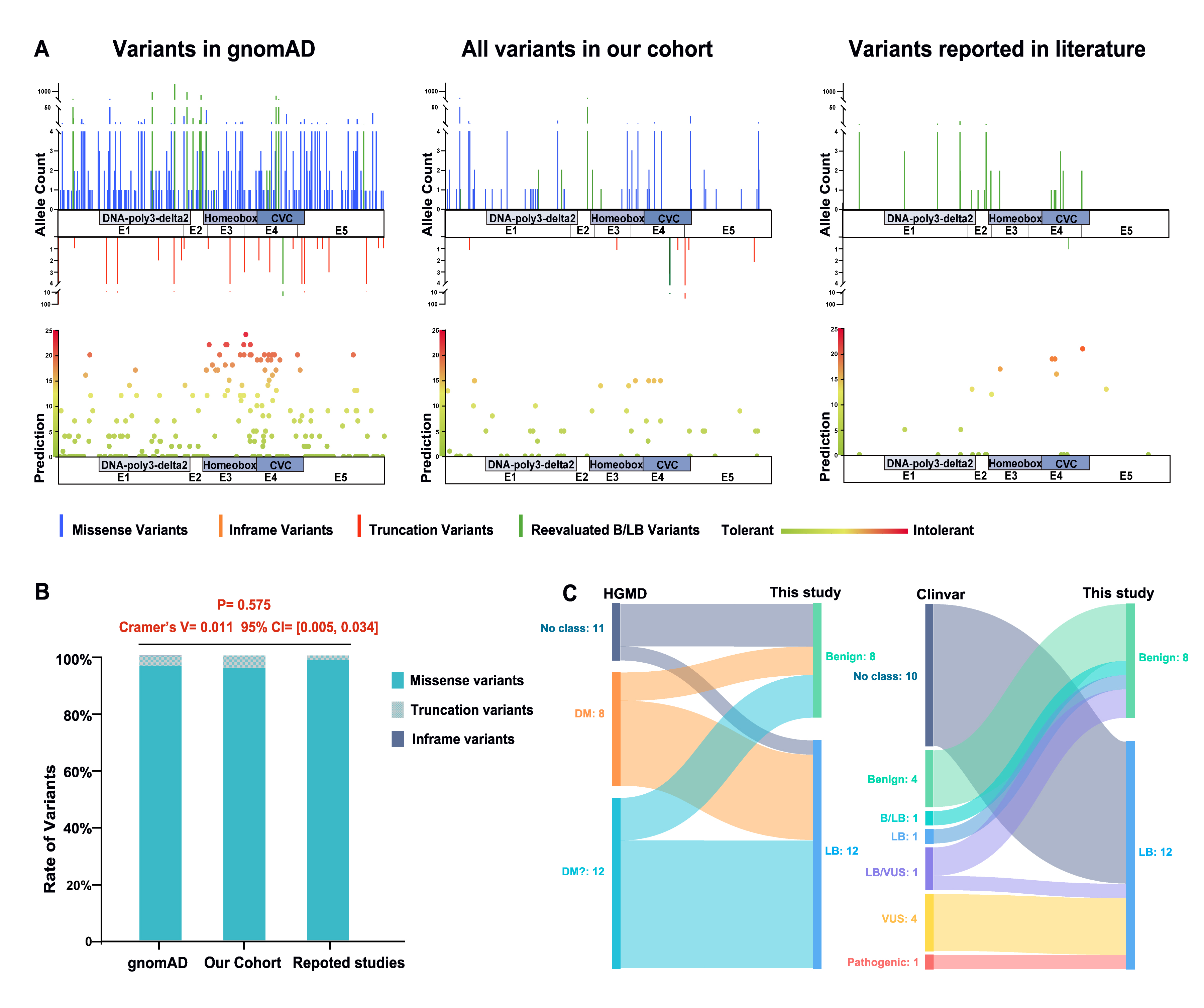
 Figure 3 of
Zhu, Mol Vis 2026; 32:70-82.
Figure 3 of
Zhu, Mol Vis 2026; 32:70-82.  Figure 3 of
Zhu, Mol Vis 2026; 32:70-82.
Figure 3 of
Zhu, Mol Vis 2026; 32:70-82. 