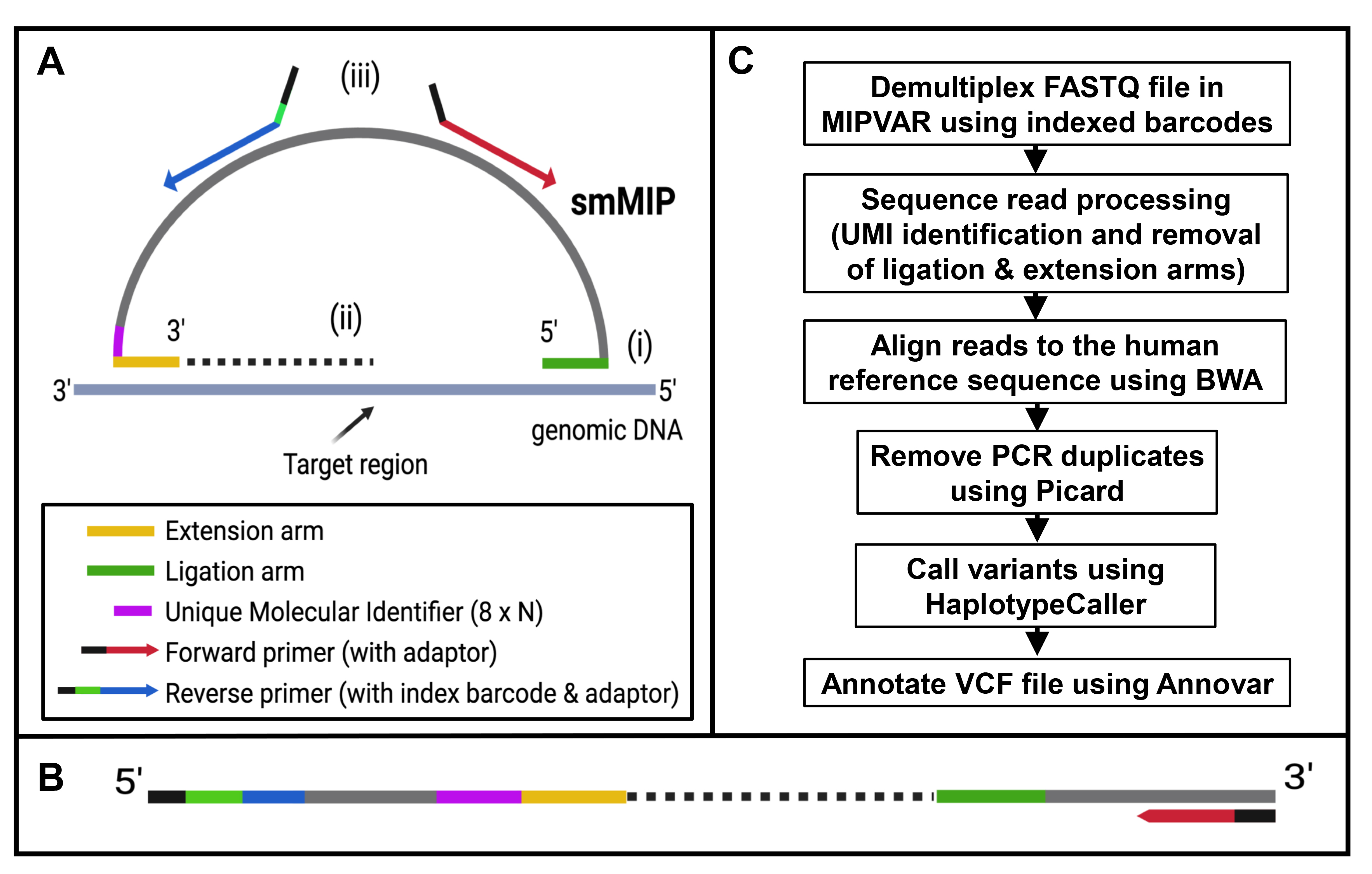
Figure 1. Summary of smMIP-based targeted sequencing workflow.
A. Schematic of one smMIP spanning target region of interest in genomic DNA. The 5′ and 3′ orientations are shown. Each smMIP
of approximately 83 nucleotides in length consists of an oligonucleotide backbone (light gray, 30 bp) with specific sequences
for primer binding (red and blue), extension (yellow, 16–22 bp), and ligation (in green, 22–28 bp) arms and the unique molecular
identifier (UMI; purple, 8 bp). The smMIP hybridizes to the target region of interest (i), then undergoes extension and ligation
to complete a DNA circle (ii). This is linearized following PCR with universal primers containing an 8-bp index barcode unique
to that sample and adaptor sequences specific to the sequencing platform being used (iii), to give the linearized product
shown in (
B). For each patient sample, a megapool of smMIPs can be used simultaneously to create a library with its own unique index,
which can be pooled with other libraries and sequenced.
C. Pipeline of data analysis following sequencing. Panels A and B were created in
BioRender. Alayed, B. (2025).
 Figure 1 of
Alayed, Mol Vis 2025; 31:486-500.
Figure 1 of
Alayed, Mol Vis 2025; 31:486-500.  Figure 1 of
Alayed, Mol Vis 2025; 31:486-500.
Figure 1 of
Alayed, Mol Vis 2025; 31:486-500.