Figure 6. Predictive outcome of PDE6A splice acceptor site variations on the exon splicing mechanism. A: Ideograms illustrate the skipping of exon 11 due to a splice acceptor site mutation at position −2 of intron 10: c.1408–2A>G.
B–D: Ideograms illustrate the predictive splicing patterns of exon 17 due to the splice acceptor site mutation at position −1
of intron 16: c.2028–1G>A. B: The prediction of a strong consensus value (CV) of 82.23 for the wild-type splice acceptor site (c.2028–1G). C: A possible scenario that would result in the skipping of exon 17 during mRNA processing. D: Predicting the creation of a new cryptic splice acceptor site (CV=74.31) with predicted deviation (∆CV) of +63.81% that
would result in the premature stop codon (p.K677Rfs24*) and eventually would lead to a truncated protein. CV=consensus values;
∆CV=reduction in consensus values.
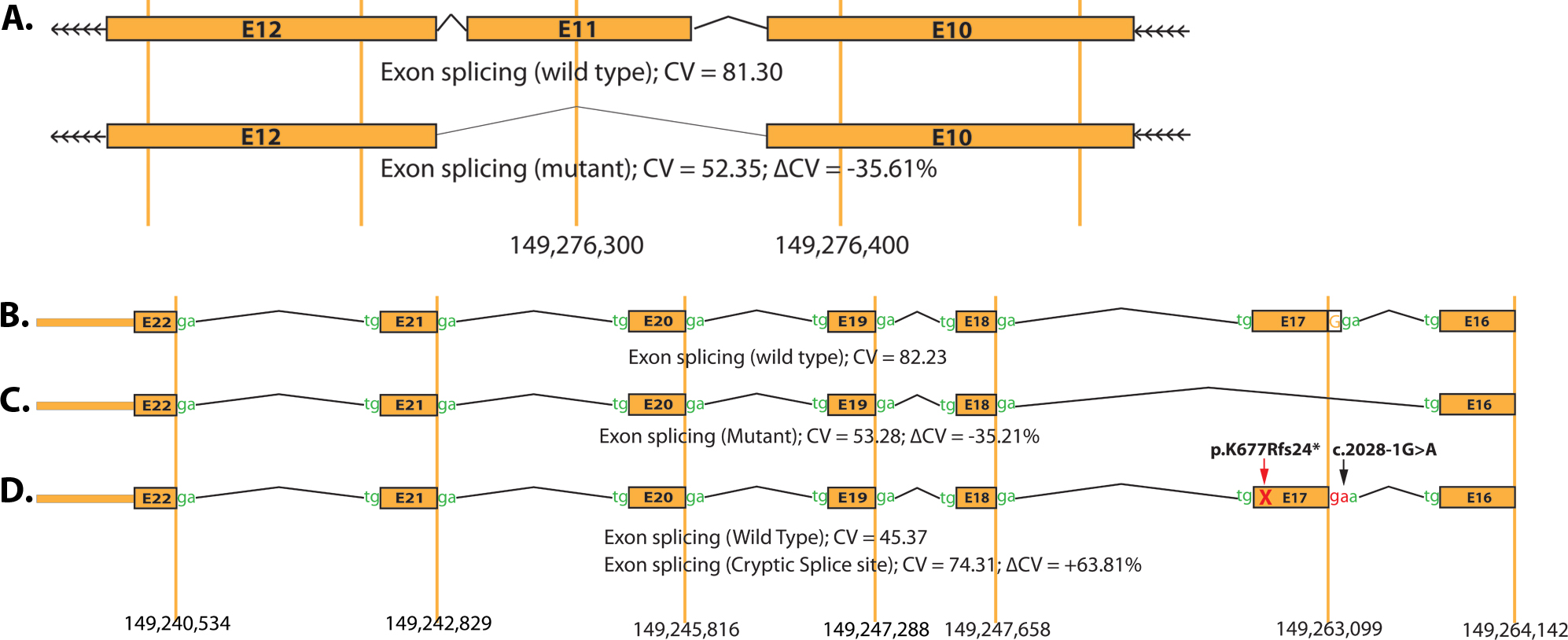
 Figure 6 of
Khan, Mol Vis 2015; 21:871-882.
Figure 6 of
Khan, Mol Vis 2015; 21:871-882.  Figure 6 of
Khan, Mol Vis 2015; 21:871-882.
Figure 6 of
Khan, Mol Vis 2015; 21:871-882. 