Figure 5. Three-dimensional domain
architecture of the tubby domain of TULP1 wild-type and mutant
proteins. A: Preferred predicted secondary structure of
normal tubby-like protein 1 (TULP1) with Thr380 and Arg482
indicated in green. In yellow, the inositol triphosphate
molecules that are predicted to bind the tubby domain of TULP1.
B: Predicted structure of part of the p.Thr380Ala mutant
protein in affected individuals of family A. The smaller size of
the alanine residue may lead to rearrangements of surrounding
residues and thereby affect putative inositol triphosphate
binding. C: Part of the predicted structure of the
p.Arg482Gln mutant protein found in affected individuals of
family B. The p.Arg482Gln variant changes a positively charged
amino acid (arginine) to a neutral residue (glutamine), which
leads to loss of interactions with two negatively charged
residues in its vicinity. Wild-type interactions are indicated
with yellow blocks.
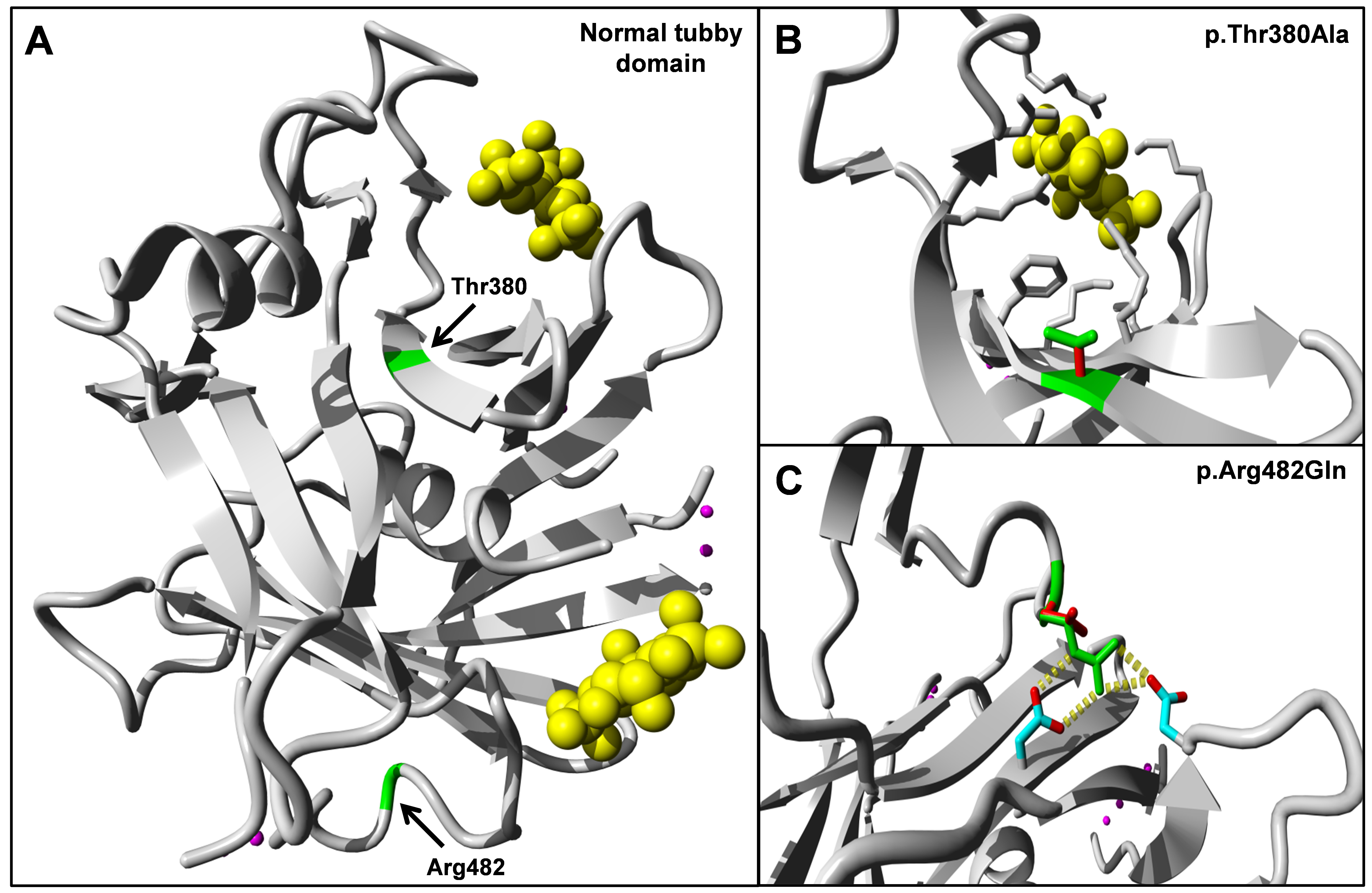
 Figure 5
of Ajmal, Mol Vis 2012; 18:1226-1237.
Figure 5
of Ajmal, Mol Vis 2012; 18:1226-1237.  Figure 5
of Ajmal, Mol Vis 2012; 18:1226-1237.
Figure 5
of Ajmal, Mol Vis 2012; 18:1226-1237. 