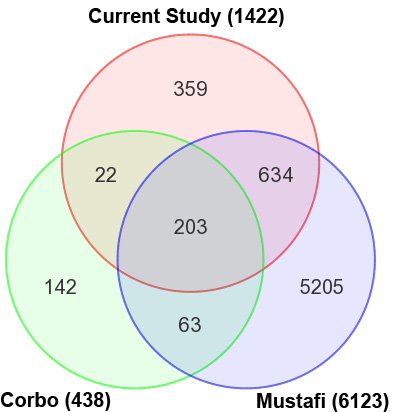
Figure 6. Comparison of the current
and previous data sets of differentially expressed transcripts
of
Nrl−/− versus wild type (WT) mouse retina.
Overlap between the differentially expressed transcripts (DETs)
identified in the current study and previous studies using mouse
retinas from the same age and genotype was determined using the
Mouse Genome Informatics (MGI) gene symbol as the identifier.
The current study includes all mRNA transcripts identified with
the Burrows-Wheeler Aligner (BWA) workflow (fold change ≥1.5 and
p value <0.05). The 438 DETs of an Affymetrix microarray
study [
38]
and 6,123 DETs from another RNA-seq study [
54]
with similar criteria were used for comparison with our study.
Of the 438 DETs from the Corbo et al. [
38]
study, 157 transcripts are not significantly differentially
expressed in our data, 11 are expressed below the fragments per
kilobase of exon model per million mapped reads (FPKM) detection
threshold of 1.0, and 38 do not map to the current annotations.
Of the 6,123 DETs from the Mustafi et al. [
54]
study, 4,858 transcripts are not significantly differentially
expressed in our study, 348 are expressed below our FPKM
detection threshold of 1.0, and 62 do not map to the current
annotations.
 Figure 6
of Brooks, Mol Vis 2011; 17:3034-3054.
Figure 6
of Brooks, Mol Vis 2011; 17:3034-3054.  Figure 6
of Brooks, Mol Vis 2011; 17:3034-3054.
Figure 6
of Brooks, Mol Vis 2011; 17:3034-3054.