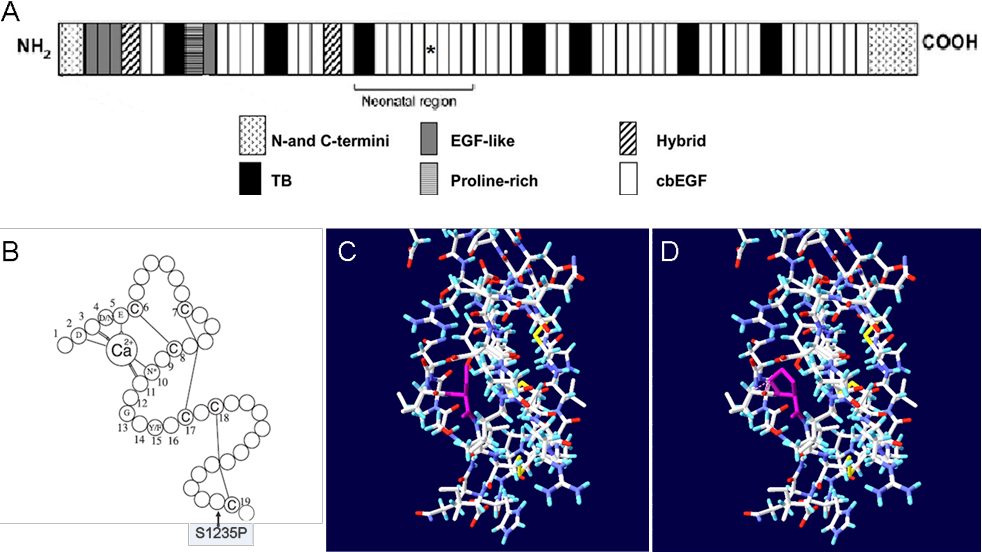
Figure 4. Structure analyses of the missense mutation in the calcium binding (cb) epidermal growth factor (EGF)-like#15 domain.
A displays the location of the affected module in the neonatal region of the fibrillin-1 domain structure.
B displays the consensus secondary structure of a prototypical cb EGF-like domain. Calcium binding in the NH
2-terminal region of the wild-type domain is mediated by the consensus sequence (D/N)-X-(D/N)(E/Q)Xm(D/N)Xn(Y/F) (m and n are
variables), and highly conserved amino acids are identified by their single-letter amino acid code. Letter C in the schematic
represents the highly conserved cysteines of cb EGF-like domain, and the lines between cysteines represent disulfide bridges.
The mutation S1235P is located at the 39th residue of the domain, which neither interferes with calcium binding of the domain
nor influences disulfide bond formation.
C displays the 3D structure of the cb EGF-like#15 domain, which is created based on the Protein data bank (PDB) template 1LMJ
(45% sequence identity) by
DeepView v4.0.1. The yellow lines represent disulfide bonds, and the purple residue represents the unaffected serine.
D displays the potential consequence of the mutation. The purple residue represents the substitute proline, and the purple
dashed lines mean steric clash with surroundings, which may lead to unstable conformation.
 Figure 4 of
Meng, Mol Vis 2011; 17:2421-2427.
Figure 4 of
Meng, Mol Vis 2011; 17:2421-2427.  Figure 4 of
Meng, Mol Vis 2011; 17:2421-2427.
Figure 4 of
Meng, Mol Vis 2011; 17:2421-2427.