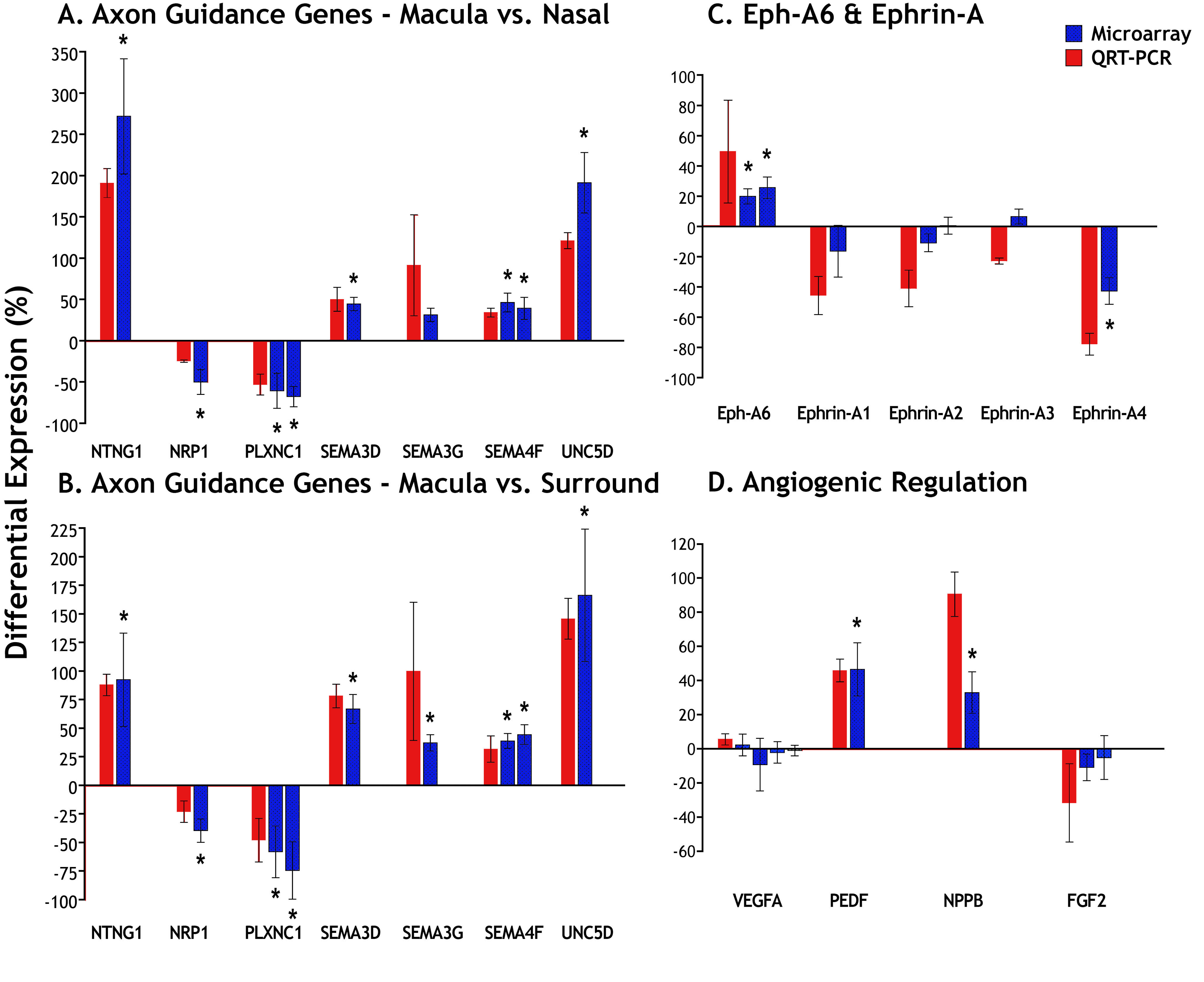
Figure 2. Measures of differential gene
expression. Microarray data are indicated by blue bars and QRT–PCR data
by red bars. We selected differentially regulated genes (*p<0.01)
identified by the microarray, binding partners of those differentially
regulated genes, as well as five other relevant genes of interest:
SEMA3G,
FGF2, VEGF, and
ephrin-A2 and
-A3.
A,
B:
Selected axonal guidance genes. These genes were significantly
upregulated or downregulated in the macula compared to either the nasal
(
A) or surround (
B) samples, with the exception of
SEMA3G,
which was significantly upregulated only in the macula versus surround
comparison (
B).
C: QRT–PCR indicates strong differential
regulation of
Eph-A6 and its ligands
ephrin-A1 and -A4
in the macula compared with nasal samples. The QRT–PCR results showed
stronger trends in differential expression compared to the microarray,
with the exception of
ephrin-A3, which QRT–PCR suggests is not
differentially expressed. Similar data were obtained from macula versus
surround comparisons.
D: The QRT–PCR confirmed significant
upregulation of
PEDF and
NPPB in the macula compared
with the nasal sample. Both the microarray and QRT–PCR findings
indicate no differential regulation of
VEGFA, the major isoform
expressed in the retina. The small downregulation of
FGF2 in
the microarray and QRT–PCR data are consistent with our previous
findings that show a downregulation of
FGF2 in macular cones,
but not in other layers of the retina, by in situ hybridization [
32].
Note that the
standard error of the microarray fold changes is derived from the
distribution of fold changes measured across the four specimens.
Abbreviations: fibroblast growth factor 2 (
FGF2); natriuretic
peptide precursor B (
NPPB); neuropilin 1 (
NRP1); netrin
G1 (
NTNG1); pigment epithelium derived factor (
PEDF);
plexin C1 (
PLXNC1); semaphorin 3D (
SEMA3D); semaphorin 3G
(
SEMA3G); semaphorin 4F (
SEMA4F); unc-5 homolog D (
C.
elegans;
UNC5D); vascular endothelial growth factor A (
VEGFA).
 Figure 2 of Kozulin, Mol Vis 2009; 15:45-59.
Figure 2 of Kozulin, Mol Vis 2009; 15:45-59.  Figure 2 of Kozulin, Mol Vis 2009; 15:45-59.
Figure 2 of Kozulin, Mol Vis 2009; 15:45-59.