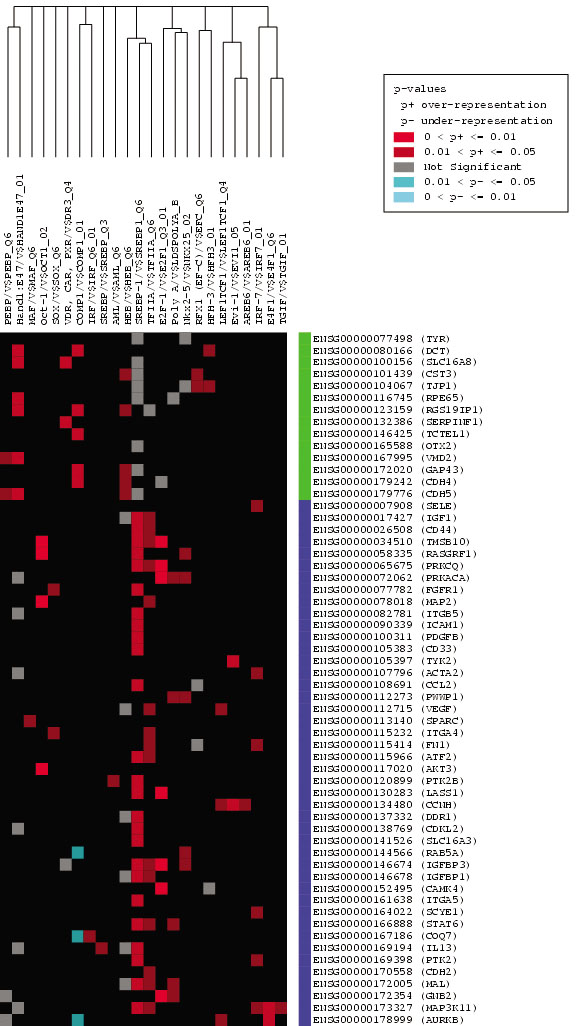
Figure 1. Candidate interaction matrix for
statistically enriched transcription response elements from promoter
analysis and interaction network toolset analysis of human gene
promoters. The retinal pigment epithelium (RPE) gene set was analyzed
by promoter analysis and interaction network toolset (PAINT) and a
graphic candidate interaction matrix (CIM) was generated as described
in Methods. The y-axis lists the Ensembl Gene identifiers for each gene
and the x-axis lists the TRANSFAC identifiers for each transcription
response element (TRE) found at least once in the promoter region of
one or more genes. Genes listed along the y-axis are divided into two
clusters that are either upregulated (blue) or down-regulated (green)
during epithelial-mesenchymal transformation (EMT) of RPE cells. TREs
listed along the x-axis are clustered according to related occurrence
pattern calculated using Jaccard's coefficient. The elements within the
matrix are color-coded based on the p-value of each TRE found in the
regulatory regions of the genes. A red dot represents a TRE that is
statistically significant and therefore over-represented in our gene
set, while a blue dot signifies an under-represented TRE and a gray dot
stands for a TRE with no statistical significance in our gene list.
This figure represents the subset of enriched TREs for the human
genome; the full CIMs for human and other genomes analyzed are shown in
Appendix 4,
Appendix 5,
Appendix 12,
and
Appendix
13.
![]() Figure 1 of Pratt,
Mol Vis 2008; 14:1414-1428.
Figure 1 of Pratt,
Mol Vis 2008; 14:1414-1428.