![]() Figure 1 of
Wimplinger, Mol Vis 2007;
13:1475-1482.
Figure 1 of
Wimplinger, Mol Vis 2007;
13:1475-1482.
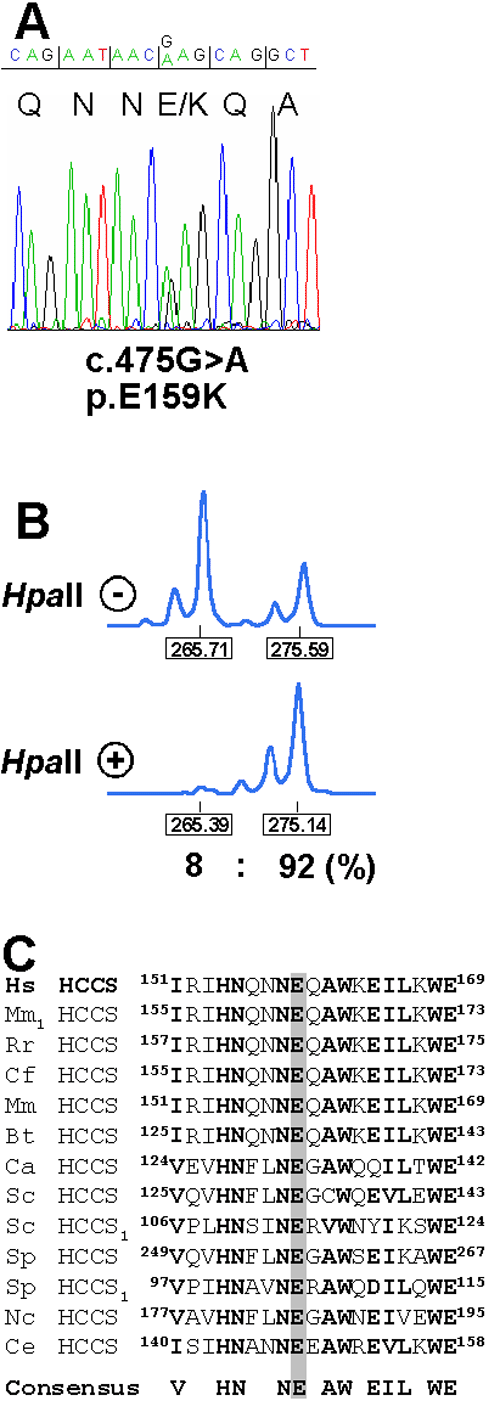
Figure 1. An HCCS missense mutation in a female with nonsyndromic microphthalmia
A: Sequence electropherogram of part of HCCS exon 5 from genomic DNA of the proband. Triplets and encoded amino acids are indicated. The patient is heterozygous for the c.475G>A mutation (p.E159K) in exon 5. B: X chromosome inactivation determined by amplification of an AR sequence polymorphism and digestion of genomic DNA isolated from lymphocytes with HpaII (indicated with + or -). The allele sizes and ratio of the X-inactivation pattern are given below the peaks. C: Partial amino acid sequence alignment of the first heme lyase targeting motif from various species. The position of amino acids is given. Evolutionary conserved residues are presented in bold. The invariant glutamate at position 159 is shaded in gray. The consensus sequence is indicated below the amino acid alignment. HCCS1 indicates specificity of this heme lyase for cytochrome c1. Hs, Homo sapiens; Mm1, Mus musculus; Rr, Rattus norvegicus; Cf, Canis familiaris; Mm, Macaca mulatta; Bt, Bos taurus; Ca, Candida albicans; Sc, S. cerevisiae; Sp, Schizosaccharomyces pombe; Nc, Neurospora crassa; Ce, Caenorhabditis elegans.