![]() Figure 2 of
Kraft, Mol Vis 2005;
11:1246-1256.
Figure 2 of
Kraft, Mol Vis 2005;
11:1246-1256.
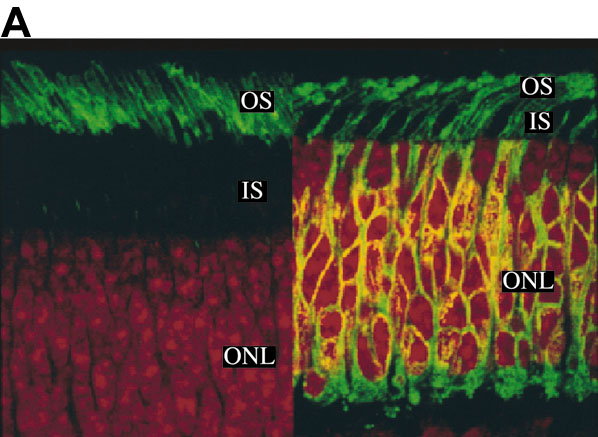
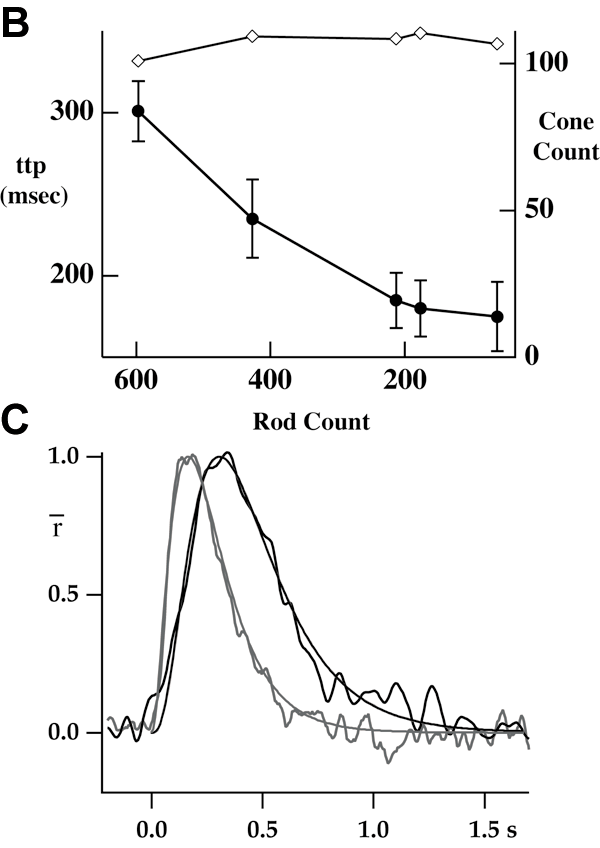
Figure 2. Rod counts predict physiological changes
Several tissue samples were analyzed for the number of rods remaining and compared to cell physiology. A: The green signal is from FITC anti-rhodopsin and the orange is from nuclear stain propidium iodide. Left panel shows normal retina in which rhodospin was restricted mainly to outer segments (OS); Right panels is from a transgenic pig and shows delocalization of rhodopsin to inner segments (IS), plasma membrane of outer nuclear layer (ONL), and synaptic region. The "ring" around a nucleus indicated a rod rather than the nuclei without the "ring", located near the top edge of the ONL that were counted as cones. In other sections, an antibody against cone transducinγ was used to stain the cones, allowing their counting directly with "positive" antibody staining. B: The level of retinal degeneration as measured by the cell counts of the surviving rods is inversely related to the time-to-peak (ttp) of the rod response to brief flashes of light (filled circles; mean±SEM, left axis). Each point represents the data for one animal; from left to right the rod count declines representing more advanced stages of degeneration. The number of cone nuclei was also counted and was apparently unaffected by the loss of rods (open diamonds, right axis). C: Dim flash responses recorded from rods isolated from a wild type animal (black trace) or a transgenic animal with an advanced state of retinal degeneration (grey trace). For wild type, the stimulus was 500 nm, 1.02 photons/μm2, average of 100 traces. For transgenic, the stimulus was 20.4 photons/μm2, average of 35 sorted "single" responses; see text for details. The smooth curves drawn through each photocurrent response are simple multistage filter models of the photoresponse, fit with three independent stages [59] and a time to peak of 165 ms (transgenic) or 310 ms (wild type).