![]() Figure 4 of
Boehlke, Mol Vis 2004;
10:867-873.
Figure 4 of
Boehlke, Mol Vis 2004;
10:867-873.
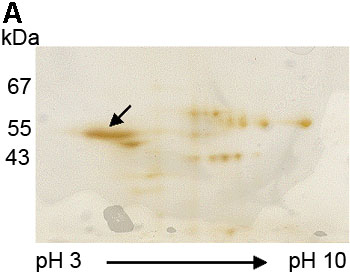
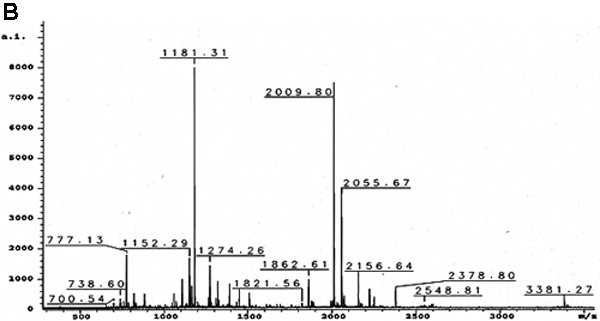
Figure 4. Confirmation of the 55 kDa protein reactive with E17 to be cytokeratin 12 by in-gel digestion and MALDI-TOF
A: The 55 kDa protein band (arrow) was excised from the silver stained, two dimensional electrophoresis gel. B: MALDI-TOF MS analysis showed a monoisotopic peptide mass fingerprint spectrum of the protein extracted from the 55 kDa band. C: A Mascot search showed the matched peptides (in red) from the isolated 55 kDa protein accounted for 51% of the protein sequence of cytokeratin 12. D: A probability based Mowse score of 147 (arrow) indicated that the identified peptides match the cytokeratin 12 protein sequence with a high degree of certainty. The shaded area represents scores with the greatest statistical uncertainty. The Mowse score is derived from (-10)log10(P), where P is the probability that the observed match is a random event. Mowse scores greater than 63 (or p<0.05) are considered a significant match of the proteins. The shaded area showed that matching of these tryptic peptides by Mascot search with other protein sequences yielded Mowse scores lower than 63 and the tryptic peptides from 55 kDa did not match significantly (p>0.05) with any protein other than cytokeratin 12 (Mowse scoring is described in detail on the Matrix Science web site).
C:
1 MDLSNNTMSL SVRTPGLSRR LSSQSVIGRP RGMSASSVGS GYGGSAFGFG 51 ASCGGGFSAA SMFGSSSGFG GGSGSSMAGG LGAGYGRALG GGSFGGLGMG 101 FGGSPGGGSL GILSGNDGGL LSGSEKETMQ NLNDRLASYL DKVRALEEAN 151 TELENKIREW YETRGTGTAD ASQSDYSKYY PLIEDLRNKI ISASIGNAQL 201 LLQIDNARLA AEDFRMKYEN ELALRQGVEA DINGLRRVLD ELTLTRTDLE 251 MQIESLNEEL AYMKKNHEDE LQSFRVGGPG EVSVEMDAAP GVDLTRLLND 301 MRAQYETIAE QNRKDAEAWF IEKSGELRKE ISTNTEQLQS SKSEVTDLRR 351 AFQNLEIELQ SQLAMKKSLE DSLAEAEGDY CAQLSQVQQL ISNLEAQLLQ 401 VRADAERQNV DHQRLLNVKA RLELEIETYR RLLDGEAQGD GLEESLFVTD 451 SKSQAQSTDS SKDPTKTRKI KTVVQEMVNG EVVSSQVQEI EELM |